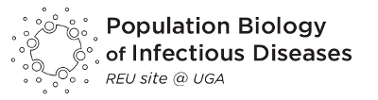Sydney Barosko, a student from Michigan State University, worked in the lab of Dr. Travis Glenn to examine pathogen diversity in local lone star ticks.
Abstract: In the world of infectious diseases, ticks play an important role as a vector in transmitting pathogens to humans, companion animals, livestock, and wildlife. Amblyomma americanum (Lone Star ticks) are known to transmit the pathogens that cause ehrlichiosis, babesiosis, Q fever, and rickettsial diseases. Few studies have been done on the pathogen diversity of Lone Star tick individuals. We used microbial 16S amplification followed by Illumina sequencing of the 16S amplicons to characterize microbial communities in A. americanum collected near Athens, GA. We examined differences in 16S sequences: 1) when two different Taq DNA polymerases were used for amplification (one with high fidelity and the other with more tolerance for low-quality samples and primer mismatches) and, 2) between 19 male and 18 female A. americanum ticks. We focused on three genera of microbes with known pathogenic strains:Â Coxiella, Rickettsia, and Ehrlichia. We did not find any significant differences between the communities when amplified with the different Taq DNA polymerases nor in the infection rate of males vs. females infected with Ehrlichia or Rickettsia. Male vs. female infection rates did, however, differ for Coxiella. The proportion of Coxiella in the microbiome was much higher in females than in males. Our work demonstrates that 16S microbiome sequencing can be an effective tool in characterizing pathogens in ticks and builds a foundation for larger-scale surveys in the future.